I was notified about this project from the 2000’s
Universally Programmable Intelligent Matter
Compare the earlier Maclennan:
Universally programmable intelligent matter (UPIM) is made from a small set of molecular building blocks that are universal in the sense that they can be rearranged to accomplish any purpose that can be described by a computer program. In effect, a computer program controls the behavior of the material at the molecular level. In some applications the molecules self-assemble a desired nanostructure by “computing” the structure and then becoming inactive). In other applications the material remains active so that it can respond, at the molecular level, to its environment or to other external conditions. An extreme case is when programmable supra-molecular clusters act as autonomous agents to achieve some end.
… with the later Buliga:
Define a molecular computer as one molecule which transforms, by random chemical reactions mediated by a collection of enzymes, into a predictable other molecule, such that the output molecule can be conceived as the result of a computation encoded in the initial molecule.
Compare the earlier Maclennan:

… with the later chemSKI rewrite
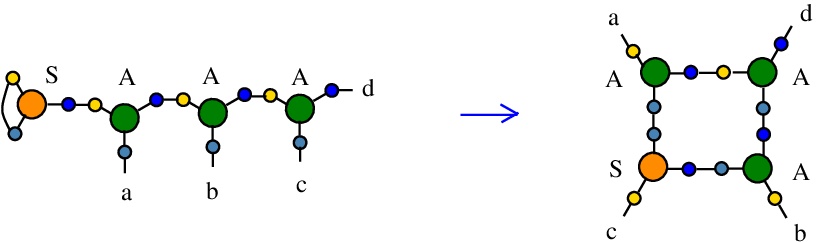
In relation to chemSKI with tokens:
- is almost the same thing! I shall update all the references, where needed.
- I bet I can easily simulate all the nanostructures from UPIM in chemSKI, which I shall do,
- and I have to go back to the chemlambda collection of simulations for chemlambda based such structures.
- I shall update Molecular computers with interaction combinators like graph rewriting systems (whenever I shall be able to)
- I return to UPIM project the challenge to: “Compute ackermann(2.2) or ackermann(2,3) with real chemistry. Or why not ackermann(3,2)? or ackermann(4,4) to see what an ackermann goo looks like, macroscopically.”
Oh I’m so relieved that I was not the first to dream about such things.